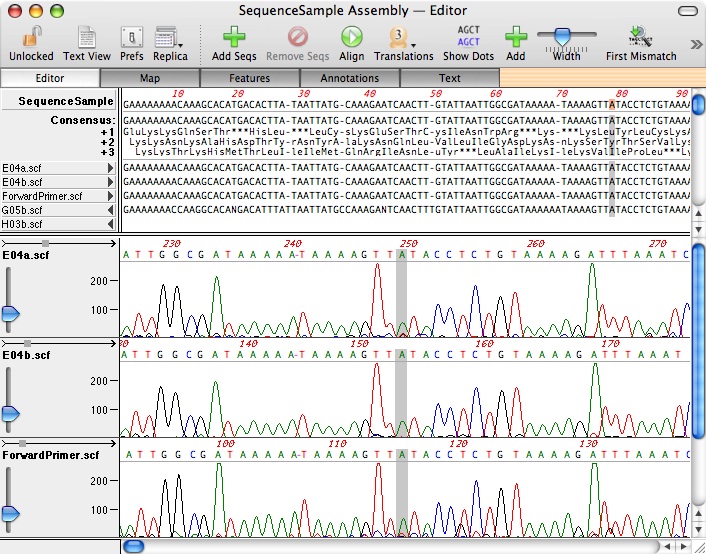
In wheat, atp6, atp8, rrn5, rrn18, rrn26, trnQ, trnK, trnfM, trnD, and trnP have multiple copies because they are in the 11 repeats of wheat mt genome (7 large and 4 short). Genes in these larger repeats or short repeats (100–1000 bp) may have multiple copies. The large repeats (>1 kb) can be divided into direct repeats and inverted repeats, the former accounting for a larger proportion of the total. 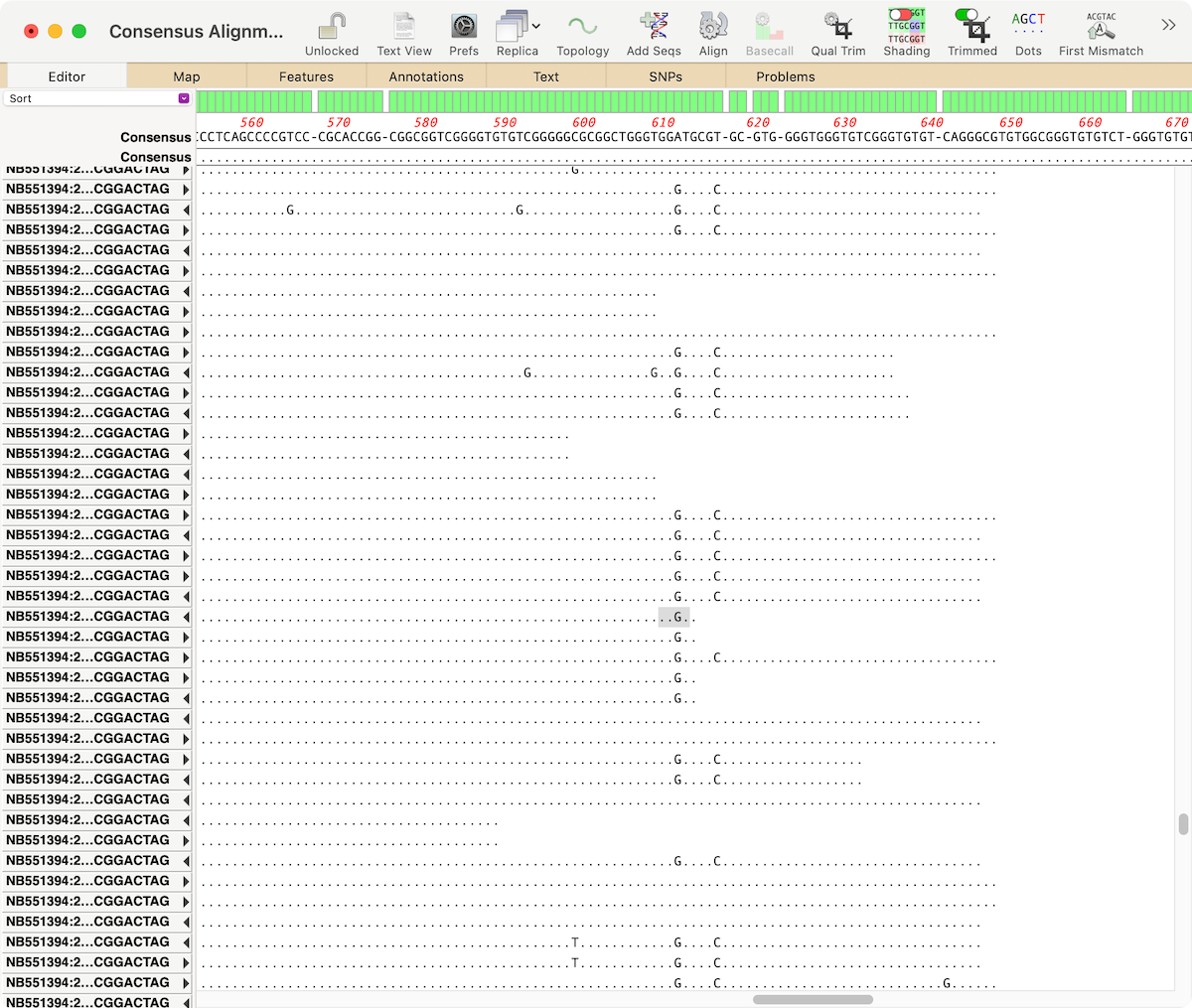
The mt genome of Cucurbita pepo contains tens of thousands of short repeats (total 371 kb), resulting in a nearly 1 Mb genome. Short repeats, usually ranging from dozens to several hundred bp, play vital roles in the evolution of plant mt genomes and may be responsible for structural variations and variable sizes in higher plant mt genomes. Methylated sites in maize mt genomes are related to tandem repeats. Many experiments showed that mitochondrial repeats contain a lot of genetic information and are also vital components of intramolecular recombination. Plant mt genomes are rich in repeats, including tandem repeats, short repeats, and large repeats. For example, the cp genomes of rice, maize, and wheat share almost the same gene orders but their mt genomes are completely different, further reflecting the complexity of plant mt genomes. While plant cp genomes have very conservative gene order, mt genomes often do not. Additionally, some mt genomes are linear molecules, such as Oryza sativa ssp. Most mt genomes are a major circular molecule but Cucumis sativus has three circular molecules. The structures of plant mt genomes can be quite complex. The variable sizes of plant mt genomes may be due to expansion of noncoding sequence and duplication of a large segment. However, the functional genes in these genomes are quite conservative. In seed plants, the size of the mt genome is highly variable, ranging from 208 Kb in Brassica hirta to 11.3 Mb in Silene conica. Plant mt genomes, encoding the main manufacturers of cellular ATP, play vital roles in the regulation of cellular metabolism. In contrast to the conserved structures of cp genomes, mt genomes are specific to each plant in their variable sizes, complex structures, multiple RNA-editing processes, frequent reorganizations, and gene loss during evolution. Due to their smaller sizes and conserved structures, the complete cp genome sequences were determined more frequently than mt genomes. With the development of next-sequencing technology, a dramatic increase in sequences of complete organelle genomes has been witnessed in the past several years. The nuclear genome carries the overwhelming majority of information, but the chloroplast and mitochondrial genomes are nonetheless indispensable as well. IntroductionĮukaryotes have three genomes, one nucleus and two organelles. raimondii mt genome may provide a crucial foundation for evolutionary analysis, molecular biology, and cytoplasmic male sterility in cotton and other higher plants. barbadense) than other rosids, and the clade formed by two Gossypium species is sister to Brassicales. raimondii, as a member of Malvaceae, is much closer to another cotton ( G. 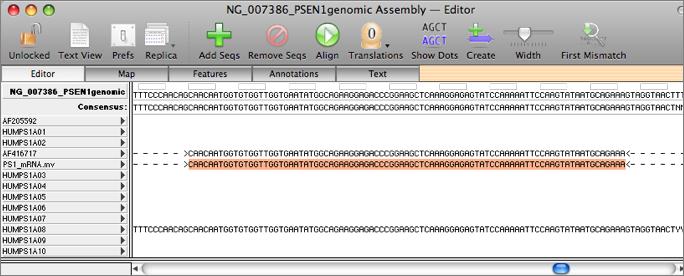
raimondii mt genome has been transferred to nucleus on Chr1, and the transfer event must be very recent. We also identified four larger repeats (63.9 kb, 10.6 kb, 9.1 kb, and 2.5 kb) in this mt genome, which may be active in intramolecular recombination in the evolution of cotton. The genome contains 39 protein-coding genes, 6 rRNA genes, and 25 tRNA genes. 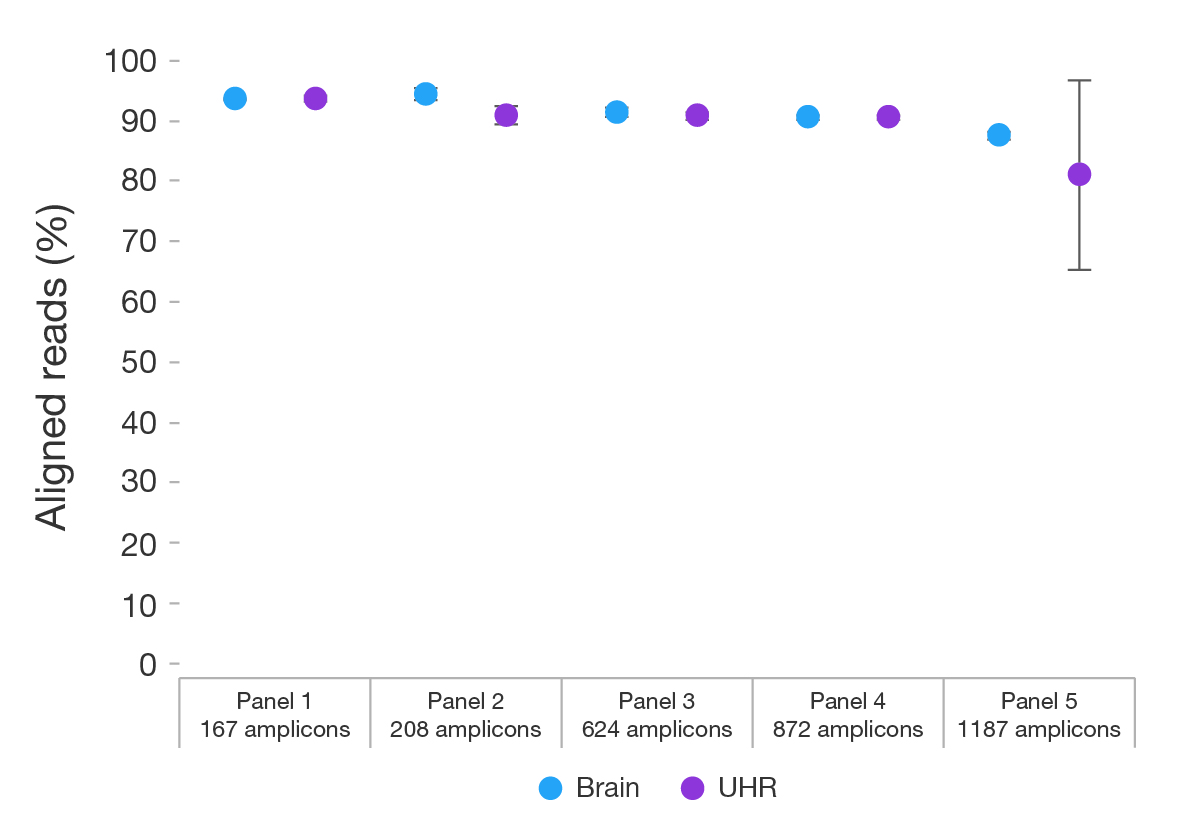
raimondii into a circular genome of length of 676,078 bp and performed comparative analyses with other higher plants. Here, we assembled the complete mitochondrial (mt) DNA sequence of G. raimondii are already available but not mitochondria. The complete nuclear and chloroplast (cp) genome sequences of G. Cotton is one of the most important economic crops and the primary source of natural fiber and is an important protein source for animal feed.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
